Oncology Nurses’ Knowledge of Pharmacogenomics Before and After Implementation of an Education Module
Objectives: To assess the efficacy of an interactive continuing education module in improving knowledge of pharmacogenomics in oncology nursing practice.
Sample & Setting: 434 inpatient and outpatient oncology nurses from a large teaching hospital in Florida and oncology nurses who practice in North Carolina.
Methods & Variables: An interactive continuing education module was created based on key information elicited from a focus group of inpatient and outpatient oncology nurses regarding their lack of knowledge on pharmacogenomics. A pre-/post-test design was implemented. Purposive sampling of oncology nurses was used.
Results: The mean pretest score was 72.7 and the post-test score was 85.9. A statistically significant difference was found between these scores. No difference in scores were found between the oncology nurses employed at urban hospitals compared to nurses at community hospitals or outpatient settings.
Implications for Nursing: Educational opportunities for pharmacogenomics should be threaded throughout nursing competencies. The continuing education module in the current article has been shown to significantly improve oncology nurses’ knowledge of genomic and pharmacogenomic information.
Jump to a section
Precision medicine has become a new innovation in cancer treatment and is growing exponentially (Das, 2017). Precision medicine is an evolving approach for disease prevention and treatment that considers variations in genes, environment, and lifestyle (Dodson, 2017). Pharmacogenomics, a subset of precision medicine, is the fastest growing specialty in pharmaceuticals. Interest in this field has risen from about 10% in 2010 to 70% in 2015, as evidenced by a poll taken by St. John’s University College of Pharmacy and Health Sciences (Ramnarain, 2016). Pharmacogenomics is “the analysis of how a person’s response to a particular drug is based on their genes” (Dodson, 2017, p. 739). This field combines pharmacology and genomics to develop safe and effective medications, along with proper dosages that should be customized to variations in a person’s genes.
Despite growth within the field of pharmacogenomics, healthcare providers have been plagued by limited knowledge on the subject, which has been identified as a major cause of the lack of pharmacogenomics use in practice (Schwartz & Issa, 2017). In addition, a lack of capability to interpret and use pharmacogenomics information was also highlighted as a barrier for the use of precision medicine in practice (Rohrer Vitek et al., 2017; Schwartz & Issa, 2017). According to Bresnick (2016), 29% of healthcare providers use precision medicine in their treatment decisions. In addition, the American Association of Colleges of Nursing and the National Institutes of Health have identified the need for nurses to take part in genetic healthcare services (Calzone et al., 2013; Calzone, Jenkins, Prows, & Masny, 2011); however, inconsistent training and education in genetics permeates the field of nongenetic clinicians (Calzone, Jenkins, Culp, Caskey, & Badzek, 2014; Cheek, Bahsore, & Brazeau, 2015; Dodson, 2014; Hoffman et al., 2016; Riddle, Gregoski, Baker, Dumas, & Jenkins, 2016).
Key findings from several studies highlight the lack of knowledge among oncology nurses, as well as the strong desire to learn more about this emerging field (Dodson, 2014; Hoffman et al., 2016; Riddle et al., 2016). A study conducted by Dodson (2014) revealed the need for increasing pharmacogenomics knowledge among nurses. Cheek et al. (2015) noted the need for education about pharmacogenomics because nurses are in key positions to ensure optimization of dosages and a reduction in adverse drug reactions. In Dodson’s (2015) study of oncology nurses’ attitudes toward pharmacogenomics, the desire to learn more about pharmacogenomics and its relevance to nursing practice was clearly documented. Based on the attitude of these nurses and the lack of knowledge surrounding pharmacogenomics, a clear need for education on this topic is imperative.
Several web-based education modules have been created for acquiring genetic and genomics competency (Calzone, Jerome-D’Emilia, et al., 2011; Global Genetics and Genomics Community, n.d). However, a pharmacogenomics educational module specific to the oncology nurses’ role could not be found through an extensive search of the literature and Internet. Therefore, an interactive continuing education pharmacogenomics module was created. The purposes of this study were to assess the efficacy of the interactive continuing education module on the role of pharmacogenomics in oncology nursing practice and to improve nurses’ knowledge of the topic.
Methods
Rogers’ diffusion of innovation theory provided the framework for examining the process of adoption of pharmacogenomics testing among oncology nurses (Rogers, 2003). Adoption of an innovation is a decision-making process in which the individual first passes from initial knowledge of the innovation to forming an attitude about this innovation. This leads to the successful adoption or rejection of the innovation. Therefore, sufficient knowledge must be provided for the individual to successfully adopt the new practice.
The first objective of this study was to create a continuing education module on pharmacogenomics that encompasses useful information for oncology nurses with the assistance of focus groups of practicing oncology nurses. The second objective was to increase nurses’ knowledge using the module related to pharmacogenomics and analyze the statistical difference between the pre- and post-test scores following use of the continuing education module. Finally, an analysis of the statistical association between work setting and knowledge was conducted.
Sample and Setting
A purposive sampling of oncology nurses was used. A sample of 32 oncology nurses from a large teaching hospital in Florida, as well as 402 oncology nurses who practice within North Carolina, were included within the study. The two populations were compared because they were easily accessible to the investigator. The sampling frame for the oncology nurses in North Carolina included all current RNs in an oncology setting within the state of North Carolina with an email address on file with the North Carolina Board of Nursing. A comprehensive list of all nurses who met these criteria was obtained from the North Carolina Board of Nursing, which resulted in 402 nurses. The sampling frame for the oncology nurses in Florida was a purposive sample of oncology nurses who worked on the oncology unit within the teaching hospital.
Education Module
Focus group interviews with oncology nurses at Moffitt Cancer Center in Tampa, Florida, were conducted to assess needs surrounding pharmacogenomics information. The focus group included nurse administrators, clinical nurse specialists, and inpatient and outpatient oncology nurses. The focus group provided information on key aspects of genomics and pharmacogenomics testing within their practice setting that they would like to learn more about.
In addition, an extensive review of the literature and current practice surrounding pharmacogenomics were reviewed to obtain the most up-to-date and relevant information surrounding pharmacogenomics. Search terms used in PubMed and CINAHL® included pharmacogenomics, precision medicine, personalized medicine, targeted therapy, and gene-based therapy. The search was limited to peer-reviewed articles published in 2011–2016 and written in English.
Finally, pertinent information related to oncology from the Pharmacogenomics Education Program (Kuo et al., 2011) and PharmGKB was incorporated into the module. The Pharmacogenomics Education Program was created for a multitude of disciplines, including pharmacists, nurses, and genetic counselors. The program is a free, evidence-based education program. PharmGKB is a pharmacogenomics database that includes information on dosing guidelines and drug labels of clinically actionable interventions based on gene–drug responses (see Figure 1). 
The completed module included basic facts on genomics and pharmacogenomics. In addition, information on drug disposition and drug target pharmacogenetic testing and targeted cancer therapy were provided. Examples seen commonly within practice, such as the gene–drug interaction between thiopurine methyltransferase and purinethol, and targeted cancer therapy for the HER2 tumor marker, were highlighted.
An interactive concept was created within the module by providing tasks, such as matching games, videos, and puzzles, to be completed to assess knowledge immediately. The study proposal was approved by the institutional review board at Winston-Salem State University before implementation of the study. A link to an online survey was placed within the continuing education module that required completion prior to accessing the education module. This link housed a questionnaire on basic genomics and pharmacogenomics from a modified version of the Knowledge and Attitude Questionnaire About Pharmacogenomic Testing (KAQ-PGx) (Roederer, Van Riper, Valgus, Knafl, & McLeod, 2012). The KAQ-PGx was developed by a team from the University of North Carolina (UNC) Center for Genomics and Society and the UNC Institute of Pharmacogenomics and Individualized Therapy (Roederer et al., 2012). The original questionnaire has been used in two previous studies assessing oncology nurses’ knowledge of genomics and pharmacogenomics (Dodson, 2014, 2015). The reliability and validity estimates of this questionnaire have not been published; however, this instrument was constructed to operationalize concepts and test relationships between knowledge and attitude in relation to use of the innovation of pharmacogenomics based on Rogers’ diffusion of innovation theory (Dodson, 2014; Rogers, 2003). The questionnaire used within the current study was adapted with permission and included the 11 demographic questions, 2 questions assessing background in genomics education, 5 basic knowledge questions about genomics, and 5 knowledge questions about pharmacogenomics testing. The 10 knowledge questions were each worth 10 points and, therefore, the total score could range from 0–100. The utilization and attitude subscales were removed to reduce the amount of time required for nurses to complete the survey.
Pilot Testing
A panel of 10 oncology nurses from the focus group piloted the questionnaire and the continuing education module. The panel consisted of oncology nurses who had been practicing more than three years. The internal consistency of the instrument was calculated. The knowledge subscale consisted of 10 items (a = 0.81), which is highly reliable. Feedback from the panel review was used to revise the module for better operationability and user friendliness. A few broken hyperlinks were fixed, along with spelling errors. The time for completion for the module and survey was determined to be 45 minutes.
To assess knowledge of pharmacogenomics, a pre-/post-test design was implemented. After the panel review of the continuing education module, a website was created that housed the pretest assessing knowledge of pharmacogenomics, the continuing education module on pharmacogenomics in oncology, and the post-test. An invitation to participate was distributed to the sample via email with a link to the website. In addition to the mean scores for the pre- and post-test, the demographic variables of age, gender, work setting (urban hospital, community hospital, or outpatient setting), previous genomics education, and oncology certification were collected.
Univariate statistics were conducted for all demographic variables: ranges, means, medians, and standard deviations for continuous variables, and frequencies and percentages for categorical variables. The total score for the basic genomics knowledge questions, pharmacogenomics knowledge questions, and the combination of these total scores were calculated for each respondent. A paired t test was conducted on the mean total score for the pre- and post-tests. A Mann-Whitney U test was conducted to assess the difference between mean knowledge scores among oncology nurses from Florida and North Carolina. In addition, all nurses who identified as working in an urban hospital setting were compared to those who identified with a community hospital work setting or outpatient work setting. A chi-square test was conducted to assess relationships between work setting and mean knowledge scores.
Results
A total of 121 oncology nurses completed the demographic portion of the module, and 99 of those finished the pretest. Seventy-eight of the 99 nurses completed the post-test, which was the total sample used for analysis. Fifty-one nurses worked in an urban hospital and 27 worked in a community or outpatient setting. Only 27 nurses had previous pharmacogenomics education and 41 had a bachelor’s in nursing degree, 21 had an associate’s degree or diploma, and the remaining 16 had a graduate degree in nursing. Fifty nurses had oncology certification. No differences were found between the two groups (urban hospital versus community or outpatient setting) regarding previous pharmacogenomics education, highest level of education, and oncology certification (p = 0.351, p = 0.205, and p = 0.596, respectively). The similarities and differences in demographic variables can be found in Table 1. 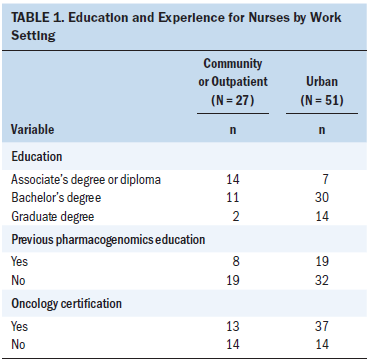
Overall, the mean pretest score was 72.7 and the post-test was 85.9 out of 100 possible points. A statistically significant difference was found between these scores (p < 0.01). The main topics that many nurses had poor knowledge of included the drug response during a patient’s lifetime and subtle differences in a patient’s genome influencing drug response. No difference in scores was found between the oncology nurses employed at urban hospitals compared to those in the community hospital or outpatient setting (p = 0.257). The scores among all the nurses in North Carolina who completed the survey were compared to the nurses in Florida to assess possible geographic differences; however, no differences in previous pharmacogenomics education, degree, and oncology certification were found between those nurses practicing in Florida compared to North Carolina (p = 0.47, p = 0.398, and p = 0.289, respectively). Finally, no difference in overall scores were found between oncology nurses in North Carolina and Florida (p = 0.898).
Discussion
Although precision medicine is growing exponentially, a lack of education exists about pharmacogenomics among oncology nurses. The results of this study match prior studies assessing knowledge of nurses as well as healthcare providers in general (Dodson, 2014; Hoffman et al., 2016; Riddle et al., 2016). Additional research is needed to determine how long the information is retained by the nurses and how it affects their patient outcomes. Based on these preliminary findings of no difference in mean score between geographic or work settings, this module may benefit many nurses in a variety of settings. The current study revealed that the continuing education module significantly improved knowledge of pharmacogenomics within the current sample; however, limitations were noted within the study. The study was not representative of the oncology nursing practice. The sample only included nurses from Florida and North Carolina, and the sampling method was different among these groups. Additional limitations included the lack of collection of number of years in nursing practice and the oncology field and the age of the nurses. These variables could have been useful to compare the characteristics of the groups at baseline. In addition, information on the reliability and validity estimates for the instrument would have improved the validity of this study. Future studies need to be replicated with a larger, representative national population with a validated tool.
Overall, this study met the first objective to create a continuing education module on pharmacogenomics. The objectives to measure the difference between the pre- and post-test scores following completion of the continuing education module and the association between work setting and knowledge were both met. However, methodologic limitations prohibit the assessment of a statistical increase in nurses’ knowledge surrounding pharmacogenomics. Future studies without these limitations would be able to test the hypothesis formulated from Rogers’ diffusion of innovation theory that increased knowledge would lead to more successful use of the innovation (Rogers, 2003).
A key aspect of this study was the inclusion of a focus group of oncology nurses to gain information on their perception of educational needs surrounding pharmacogenomics. The incorporation of these nurses added a new perspective that has not been added to other pharmacogenomics modules based on a thorough review of the literature. One of the recurring comments from the focus group was the great need for pharmacogenomics education within oncology nursing practice. Therefore, it is imperative to seek avenues for providing continuing education modules to all oncology nurses.
Implications for Nursing Practice
Information about pharmacogenomics will allow nurses to participate in research, move practice forward, and educate patients. Nurses should recognize the value of how pharmacogenomics influence their patients’ treatment and find ways to advocate for specific genetic tests and clinical trials that may be applicable to their patients’ condition. This knowledge will promote a holistic plan of care. It is important to seek avenues for providing education along with other clinical resources related to precision medicine to all oncology nurses to improve pharmacogenomics knowledge. 
Conclusion
Based on the results, educational opportunities for increasing knowledge about pharmacogenomics should be included in nursing competencies. The education module has met a need to make pharmacogenomics information relevant to nursing practice.
About the Author(s)
Crystal Dodson, PhD, RN, MSN, is an assistant professor in the School of Nursing at the University of North Carolina in Wilmington. No financial relationships to disclose. Dodson can be reached at dodsonc@uncw.edu, with copy to ONFEditor@ons.org. (Submitted October 2017. Accepted February 16, 2018.)
References
Bresnick, J. (2016). Precision medicine, data exchange, CDS on new ONC leader’s agenda. Health Analytics. Retrieved from https://healthitanalytics.com/news/precision-medicine-data-exchange-cds…
Calzone, K.A., Jenkins, J., Bakos, A.D., Cashion, A.K., Donaldson, N., Feero, W.G., . . . Webb, J.A. (2013). A blueprint for genomic nursing science. Journal of Nursing Scholarship, 45, 96–104. https://doi.org/10.1111/jnu.12007
Calzone, K.A., Jenkins, J., Culp, S., Caskey, S., & Badzek, L. (2014). Introducing a new competency into nursing practice. Journal of Nursing Regulation, 5, 40–47.
Calzone, K.A., Jenkins, J., Prows, C.A., & Masny, A. (2011). Establishing the outcome indicators for the essential nursing competencies and curricula guidelines for genetics and genomics. Journal of Professional Nursing, 27, 179–191. https://doi.org/10.1016/j.profnurs.2011.01.001
Calzone, K.A., Jerome-D’Emilia, B., Jenkins, J., Goldgar, C., Rackover, M., Jackson, J., . . . Feero, W.G. (2011). Establishment of the genetic/genomic competency center for education. Journal of Nursing Scholarship, 43, 351–358. https://doi.org/10.1111/j.1547-5069.2011.01412.x
Cheek, D.J., Bashore, L., & Brazeau, D.A. (2015). Pharmacogenomics and implications for nursing practice. Journal of Nursing Scholarship, 47, 496–504. https://doi.org/10.1111/jnu.12168
Das, R. (2017). Drug industry bets big on precision medicine: Five trends shaping care delivery. Forbes. Retrieved from https://www.forbes.com/sites/reenitadas/2017/03/08/drug-development-ind…
Dodson, C. (2014). Knowledge and attitudes of oncology nurses regarding pharmacogenomic testing [Online exclusive]. Clinical Journal of Oncology Nursing, 18, E64–E70. https://doi.org/10.1188/14.CJON.E64-E70
Dodson, C. (2015). Attitudes of oncology nurses concerning pharmacogenomics. Personalized Medicine, 12, 559–562. https://doi.org/10.2217/pme.15.37
Dodson, C.H. (2017). Pharmacogenomics: Principles and relevance to oncology nursing. Clinical Journal of Oncology Nursing, 21, 739–745. https://doi.org/10.1188/17.CJON.739-745
Global Genetics and Genomics Community. (n.d). Genomic health care simulations. Retrieved from http://genomicscases.net/en
Hoffman, S.L., Reid Kaufman, R.R., Ferrari, S., Alexander, S.A., Rosenzweig, M.Q., & Wesmiller, S.W. (2016). Beyond BRCA: A pilot program to assess and improve knowledge of pharmacogenomic testing among advanced practitioners in a breast cancer treatment setting. Journal of the Advanced Practitioner in Oncology, 7, 382–389.
Kuo, G.M., Ma, J.D., Lee, K.C., Halpert, J.R., Bourne, P.E., Ganiats, T.G., & Taylor, P. (2011). Institutional profile: University of California San Diego Pharmacogenomics Education Program (PharmGenEd™): Bridging the gap between science and practice. Pharmacogenomics, 12, 149–153. https://doi.org/10.2217/pgs.10.213
Ramnarain, R.R. (2016). Pharmacogenomics and IPhO: Growing together. Industry Pharmacists Organization. Retrieved from https://www.industrypharmacist.org/blog_post.php?id=113
Riddle, D., Gregoski, M., Baker, K., Dumas, B., & Jenkins, C.H. (2016). Impressions of pharmacogenomic testing among certified registered nurse anesthetists: A mixed-method study. Pharmacogenomics, 17, 593–602. https://doi.org/10.2217/pgs.16.3
Roederer, M.W., Van Riper, M., Valgus, J., Knafl, G., & McLeod, H. (2012). Knowledge, attitudes and education of pharmacists regarding pharmacogenetic testing. Personalized Medicine, 9, 19–27. https://doi.org/10.2217/pme.11.87
Rogers, E.M. (2003). Diffusion of innovations (5th ed.). New York, NY: The Free Press.
Rohrer Vitek, C.R., Abul-Husn, N.S., Connolly, J.J., Hartzler, A.L., Kitchner, T., Peterson, J.F., . . . Prows, C.A. (2017). Healthcare provider education to support integration of pharmacogenomics in practice: The eMERGE Network experience. Pharmacogenomics, 18, 1013–1025. https://doi.org/10.2217/pgs-2017-0038
Schwartz, E.J., & Issa, A.M. (2017). The role of hospital pharmacists in the adoption and use of pharmacogenomics and precision medicine. Personalized Medicine, 14, 27–35. https://doi.org/10.2217/pme-2016-0063




